 |
In progress |
Lindly O, Zuckerman K, Hocking TD, Folch DC, Sutherland V, Curry T |
Predict, Profile, Prioritize: Machine Learning Reveals Hidden Patterns in Autism Service Access Disparities |
Abstract submitted to INSAR 2026 conference |
Reproducible |
 |
In progress |
Agyapong D, Nguyen T, Aslam B, Propster J, Folch D, Hungate B, Marks J, Gurney K, Lindly O, Hocking TD |
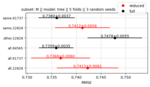
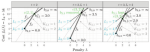
Cross-validation with subsets and downsampling for optimizing training in machine learning |
In progress |
Slides, Python Software |
 |
In progress |
Jorge E, Hocking TD, Foissac S, Neuvial P, Vialaneix N |
hicream: a flexible framework to identify significantly different regions in Hi-C data |
Under review at Bioinformatics |
Reproducible |
 |
In progress |
Agyapong D, Chiquet J, Marks J, Hocking TD |
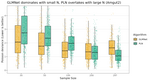
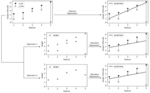
Understanding When Poisson Log-Normal Models Outperform Penalized Poisson Regression for Microbiome Count Data |
Under review at Biometrics |
Reproducible |
 |
In progress |
Nguyen TL, Hocking TD |
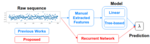
Interval Regression: A Comparative Study with Proposed Models |
Preprint arXiv:2503.02011, under review at Statistics and Computing |
Preprint, Reproducible |
 |
In progress |
Nguyen TL, Hocking TD |
Penalty Learning for Optimal Partitioning via Automatic Feature Extraction |
Preprint arXiv:2505.07413, under review at Computational Statistics |
Preprint, Reproducible |
 |
In progress |
Agyapong D, Beatty BH, Kennedy PG, Marks JC, Hocking TD |
Fused Lasso Improves Accuracy of Co-occurrence Network Inference in Grouped Samples |
Preprint arXiv:2509.09413, under review at Journal of Computational Biology |
Reproducible, Software |
 |
In progress |
Sutherland V, Hocking TD, Folch DC, Underwood Carrasco VI, Zuckerman KE, Lindly OJ |
Autism Diagnostic Determinations of Primary Care Providers from 2013 to 2023 |
Under review at Academic Pediatrics |
Related poster Trends in Autism Diagnostic Determinations by Primary Care Providers Among U.S. Children from 2013 to 2023: Effects of Child Age, Race and Ethnicity, and Poverty Level was selected as one of the Top-Rated Abstracts at INSAR 2025 conference. |
 |
In progress |
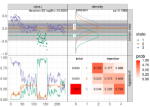
Hocking TD |
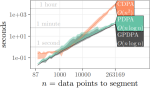
Comparing binsegRcpp with other implementations of binary segmentation for changepoint detection |
Under review at Journal of Statistical Software |
Preprint, Reproducible, Software |
 |
In progress |
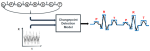
Hocking TD |
Teaching Hidden Markov Models Using Interactive Data Visualization |
In progress |
Reproducible, Slides |
 |
In progress |
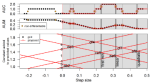
Hocking TD |
mlr3resampling for machine learning benchmarks in R |
In progress |
Software, Slides |
 |
In progress |
Fowler J, Hocking TD |
Efficient line search for optimizing Area Under the ROC Curve in gradient descent |
Preprint arXiv:2410.08635, under review at Computational Intelligence |
Preprint, Talk announcement for JSM’23 Toronto, Video of talk at Université Laval July 2023, Slides PDF, source |
 |
In progress |
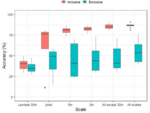
Hocking TD |
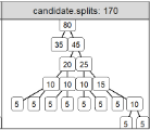
Finite Sample Complexity Analysis of Binary Segmentation |
Preprint arXiv:2410.08654, under review at Canadian Journal of Statistics |
Preprint, Reproducible, Software |
 |
In progress |
Thibault G, Morin-Bernard A, Sylvain J-D, Drolet G, Roussel J-R, Hocking TD, Achim A |
Spatial characterization of burn severity in a boreal forest using high-resolution satellite imagery |
Under review at International Journal of Applied Earth Observation and Geoinformation |
|
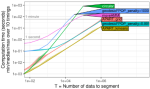 |
In progress |
Truong C, Hocking TD |
Efficient change-point detection for multivariate circular data |
Under review in JRSSC |
Reproducible, Software |
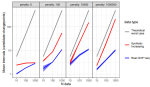 |
In progress |
Hocking TD |
Why does functional pruning yield such fast algorithms for optimal changepoint detection? |
In progress |
Reproducible, talks: semi-plenary at JDS’25 slides, TRIPODS video |
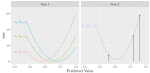 |
In progress |
Rust KR, Hocking TD |
A Log-linear Gradient Descent Algorithm for Unbalanced Binary Classification using the All Pairs Squared Hinge Loss |
Preprint arXiv:2302.11062, under review at Journal of Machine Learning Research |
Preprint |
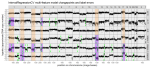 |
In progress |
Hocking TD, Killick R |
Introduction to optimal changepoint detection algorithms |
Tutorial at international useR 2017 conference, textbook in progress |
Conference, Book, Reproducible |
 |
In progress |
Hocking TD, Ekstrøm CT |
Understanding and Creating Interactive Graphics |
Tutorial at international useR 2016 conference, textbook in progress |
Conference, French Manual, English Manual, Reproducible |
 |
In progress |
Hocking TD |
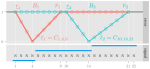
A breakpoint detection error function for segmentation model selection and validation |
Preprint arXiv:1509.00368 |
Preprint, Software, Reproducible |
 |
In progress |
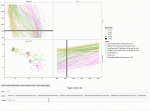
Venuto D, Hocking TD, Sphanurattana L, Sugiyama M |
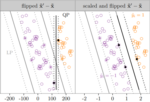
Support vector comparison machines |
Preprint arXiv:1401.8008 |
Preprint, Software, Reproducible |
 |
2026 |
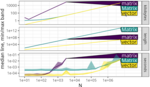
Amoakohene D, Chetia A, Steinmacher I, Hocking TD |
Asymptotic benchmarking using the atime package |
R Journal (in press) |
Reproducible, Software |
 |
2026 |
Hocking TD, Thibault G, Bodine CS, Arellano PN, Shenkin AF, Lindly O |
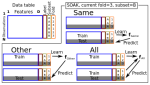
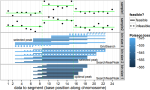
SOAK: Same/Other/All K-fold cross-validation for estimating similarity of patterns in data subsets |
Statistical Analysis and Data Mining 19(1) |
DOI, Preprint, Software, Reproducible, Local PDF |
 |
2026 |
Hocking TD |
Visualisation interactive de données dans R avec animint2 |
Université de Sherbrooke, fabriqueREL, license CC_BY-SA |
Netlify, DOI |
 |
2025 |
Oliveira P, Amoakohene D, Hocking TD, Gerosa M, Steinmacher I |
Governance Matters: Lessons from Restructuring the data.table OSS Project |
2025 IEEE International Conference on Software Maintenance and Evolution (ICSME) |
IEEE, Abstract, Preprint |
 |
2025 |
Agyapong D, Propster JR, Marks J, Hocking TD |
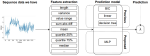
Cross-Validation for Training and Testing Co-occurrence Network Inference Algorithms |
BMC Bioinformatics 26(74) |
Publisher, Preprint, Reproducible, Local PDF |
 |
2025 |
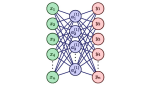
Nguyen TL, Hocking TD |
Penalty Learning for Optimal Partitioning using Multilayer Perceptron |
Statistics and Computing 35 |
Publisher, Preprint, Reproducible |
 |
2024 |
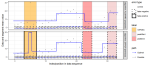
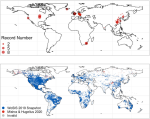
Gurney KR, Aslam B, Dass P, Gawuc L, Hocking TD, Barber JJ, Kato A |
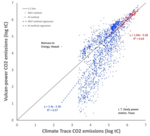
Assessment of the Climate Trace global powerplant CO2 emissions |
Environmental Research Letters 19(11) |
Publisher, Local PDF |
 |
2024 |
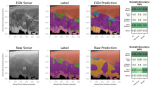
Kaufman JM, Stenberg AJ, Hocking TD |
Functional Labeled Optimal Partitioning |
Journal of Computational and Graphical Statistics 33(4) |
DOI, Preprint, Software, Reproducible, Preliminary code |
 |
2024 |
Bodine CS, Buscombe D, Hocking TD |
Automated River Substrate Mapping From Sonar Imagery With Machine Learning |
Journal of Geophysical Research: Machine Learning and Computation 1(3) |
DOI, Preprint, Software |
 |
2024 |
Tao F, Houlton BZ, Frey SD, Lehmann J, Manzoni S, Huang Y, Jiang L, Mishra U, Hungate BA, Schmidt MWI, Reichstein M, Carvalhais N, Ciais P, Wang Y-P, Ahrens B, Hugelius G, Hocking TD, Lu X, Shi Z, Viatkin K, Vargas R, Yigini Y, Omuto C, Malik AA, Peralta G, Cuevas-Corona R, Paolo LED, Luotto I, Liao C, Liang Y-S, Saynes VS, Huang X, Luo Y |
Reply to: Model uncertainty obscures major driver of soil carbon |
Nature 627(8002) |
Publisher, Preprint, Local PDF |
 |
2023 |
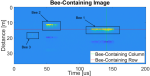
Sweeney N, Xu C, Shaw JA, Hocking TD, Whitaker BM |
Insect Identification in Pulsed Lidar Images Using Changepoint Detection Algorithms |
2023 Intermountain Engineering, Technology and Computing (IETC) |
Publisher, Local PDF |
 |
2023 |
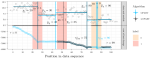
Hocking TD, Srivastava A |
Labeled optimal partitioning |
Computational Statistics 38 |
Publisher, Preprint, Video, Software, Reproducible |
 |
2023 |
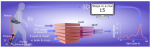
Harshe K, Williams JR, Hocking TD, Lerner ZF |
Predicting Neuromuscular Engagement to Improve Gait Training With a Robotic Ankle Exoskeleton |
IEEE Robotics and Automation Letters 8(8) |
Publisher, Local PDF |
 |
2023 |
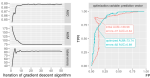
Hillman J, Hocking TD |
Optimizing ROC Curves with a Sort-Based Surrogate Loss for Binary Classification and Changepoint Detection |
Journal of Machine Learning Research 24(70) |
Publisher, Preprint, Software, Reproducible, Video, Local PDF |
 |
2023 |
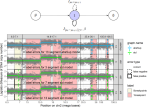
Runge V, Hocking TD, Romano G, Afghah F, Fearnhead P, Rigaill G |
gfpop: An R Package for Univariate Graph-Constrained Change-Point Detection |
Journal of Statistical Software 106(6) |
Publisher, Preprint, Software, GUI, Reproducible, Local PDF |
 |
2023 |
Tao F, Huang Y, Hungate BA, Manzoni S, Frey SD, Schmidt MWI, Reichstein M, Carvalhais N, Ciais P, Jiang L, Lehmann J, Mishra U, Hugelius G, Hocking TD, Lu X, Shi Z, Viatkin K, Vargas R, Yigini Y, Omuto C, Malik AA, Perualta G, Cuevas-Corona R, Paolo LED, Luotto I, Liao C, Liang Y-S, Saynes VS, Huang X, Luo Y |
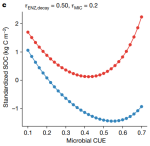
Microbial carbon use efficiency promotes global soil carbon storage |
Nature 618 |
Publisher, Local PDF |
 |
2022 |
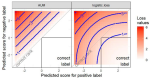
Barr JR, Hocking TD, Morton G, Thatcher T, Shaw P |
Classifying Imbalanced Data with AUM Loss |
2022 Fourth International Conference on Transdisciplinary AI (TransAI) |
Publisher, Local PDF |
 |
2022 |
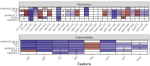
Hocking TD, Barr JR, Thatcher T |
Interpretable linear models for predicting security vulnerabilities in source code |
2022 Fourth International Conference on Transdisciplinary AI (TransAI) |
Publisher, Local PDF |
 |
2022 |
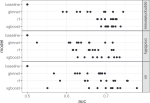
Barr JR, Shaw P, Abu-Khzam FN, Thatcher T, Hocking TD |
Graph embedding: A methodological survey |
2022 fourth international conference on transdisciplinary AI (TransAI) |
Publisher, Local PDF |
 |
2022 |
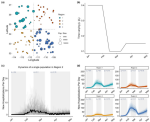
Chaves AP, Egbert J, Hocking T, Doerry E, Gerosa MA |
Chatbots Language Design: The Influence of Language Variation on User Experience with Tourist Assistant Chatbots |
ACM Trans. Comput.-Hum. Interact. 29(2) |
Publisher, Preprint, Local PDF |
 |
2022 |
Mihaljevic JR, Borkovec S, Ratnavale S, Hocking TD, Banister KE, Eppinger JE, Hepp C, Doerry E |
SPARSEMODr: Rapidly simulate spatially explicit and stochastic models of COVID-19 and other infectious diseases |
Biology Methods and Protocols 7(1) |
Publisher, Preprint, Software |
 |
2022 |
Hocking TD |
Introduction to machine learning and neural networks |
Chapter in Land Carbon Cycle Modeling: Matrix Approach, Data Assimilation, and Ecological Forecasting, edited by Yiqi Luo, published by CRC Press |
Publisher, My chapter, Reproducible, Video |
 |
2022 |
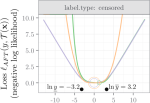
Vargovich J, Hocking TD |
Linear time dynamic programming for computing breakpoints in the regularization path of models selected from a finite set |
Journal of Computational and Graphical Statistics 31(2) |
Publisher, Preprint, Software, Reproducible, Local PDF |
 |
2022 |
Barnwal A, Cho H, Hocking T |
Survival Regression with Accelerated Failure Time Model in XGBoost |
Journal of Computational and Graphical Statistics 31(4) |
Publisher, Preprint, Software, Documentation, Video, Reproducible, Local PDF |
 |
2022 |
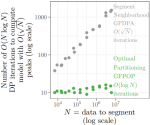
Hocking TD, Rigaill G, Fearnhead P, Bourque G |
Generalized Functional Pruning Optimal Partitioning (GFPOP) for Constrained Changepoint Detection in Genomic Data |
Journal of Statistical Software 101(10) |
Publisher, Software, Reproducible, Slides, Video |
 |
2021 |
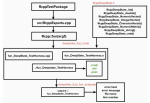
Kolla AC, Groce A, Hocking TD |
Fuzz Testing the Compiled Code in R Packages |
2021 IEEE 32nd International Symposium on Software Reliability Engineering (ISSRE) |
Publisher, Software, GitHub Action, Abstract, Video, Blog, Results, Local PDF |
 |
2021 |
Liehrmann A, Rigaill G, Hocking TD |
Increased peak detection accuracy in over-dispersed ChIP-seq data with supervised segmentation models |
BMC Bioinformatics 22(323) |
Publisher, Reproducible, Software |
 |
2021 |
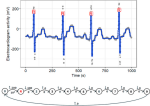
Fotoohinasab A, Hocking T, Afghah F |
A greedy graph search algorithm based on changepoint analysis for automatic QRS complex detection |
Computers in Biology and Medicine 130 |
Publisher, Preprint |
 |
2021 |
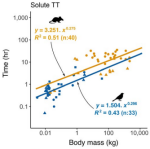
Abraham AJ, Prys-Jones TO, De Cuyper A, Ridenour C, Hempson GP, Hocking T, Clauss M, Doughty CE |
Improved estimation of gut passage time considerably affects trait-based dispersal models |
Functional Ecology 35(4) |
Publisher |
 |
2021 |
Hocking TD |
Wide-to-tall Data Reshaping Using Regular Expressions and the nc Package |
The R Journal 13(1) |
Publisher, Software, Reproducible |
 |
2020 |
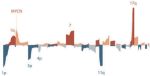
Fotoohinasab A, Hocking T, Afghah F |
A Graph-constrained Changepoint Detection Approach for ECG Segmentation |
2020 42nd Annual International Conference of the IEEE Engineering in Medicine Biology Society (EMBC) |
Publisher, Local PDF |
 |
2020 |
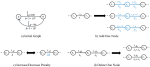
Fotoohinasab A, Hocking T, Afghah F |
A Graph-Constrained Changepoint Learning Approach for Automatic QRS-Complex Detection |
2020 54th Asilomar Conference on Signals, Systems, and Computers |
Publisher, Abstract, Preprint, Local PDF |
 |
2020 |
Hocking TD, Rigaill G, Fearnhead P, Bourque G |
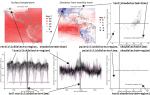
Constrained Dynamic Programming and Supervised Penalty Learning Algorithms for Peak Detection in Genomic Data |
Journal of Machine Learning Research 21(87) |
Publisher, Preprint, Software, Reproducible, Local PDF |
 |
2020 |
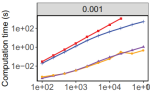
Hocking TD, Bourque G |
Machine Learning Algorithms for Simultaneous Supervised Detection of Peaks in Multiple Samples and Cell Types |
Proc. Pacific Symposium on Biocomputing |
Publisher, Software, Preprint, Reproducible |
 |
2019 |
Jewell SW, Hocking TD, Fearnhead P, Witten DM |
Fast nonconvex deconvolution of calcium imaging data |
Biostatistics 21(4) |
Pubmed |
 |
2019 |
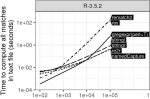
Sievert C, VanderPlas S, Cai J, Ferris K, Khan FUF, Hocking TD |
Extending ggplot2 for Linked and Animated Web Graphics |
Journal of Computational and Graphical Statistics 28(2) |
Publisher, Software, Reproducible, Interactive Figures, Local PDF |
 |
2019 |
Hocking TD |
Comparing namedCapture with other R packages for regular expressions |
The R Journal 11(2) |
Publisher, Software, Reproducible |
 |
2018 |
Depuydt P, Boeva V, Hocking TD, Cannoodt R, Ambros IM, Ambros PF, Asgharzadeh S, Attiyeh EF, Combaret Vé, Defferrari R, Fischer M, Hero B, Hogarty MD, Irwin MS, Koster J, Kreissman S, Ladenstein R, Lapouble E, Laureys Gè, London WB, Mazzocco K, Nakagawara A, Noguera R, Ohira M, Park JR, Pötschger U, Theissen J, Tonini GP, Valteau-Couanet D, Varesio L, Versteeg R, Speleman F, Maris JM, Schleiermacher G, Preter KD |
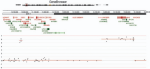
Genomic amplifications and distal 6q loss: novel markers for poor survival in high-risk neuroblastoma patients |
JNCI: Journal of the National Cancer Institute 110(10) |
Publisher |
 |
2018 |
Depuydt P, Koster J, Boeva V, Hocking TD, Speleman F, Schleiermacher G, De Preter K |
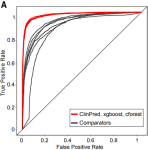
Meta-mining of copy number profiles of high-risk neuroblastoma tumors |
Scientific data 5(1) |
Publisher |
 |
2018 |
Alirezaie N, Kernohan KD, Hartley T, Majewski J, Hocking TD |
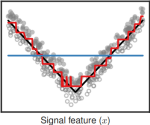
ClinPred: Prediction Tool to Identify Disease-Relevant Nonsynonymous Single-Nucleotide Variants |
The American Journal of Human Genetics 103(4) |
DOI, Software, used in dbNSFP |
 |
2017 |
Drouin A, Hocking T, Laviolette F |
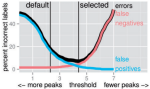
Maximum Margin Interval Trees |
Advances in Neural Information Processing Systems 30 |
Publisher, Software, Reproducible, Preprint, Video |
 |
2017 |
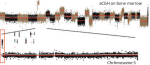
Hocking TD, Goerner-Potvin P, Morin A, Shao X, Pastinen T, Bourque G |
Optimizing ChIP-seq peak detectors using visual labels and supervised machine learning |
Bioinformatics 33(4) |
Pubmed, Software, Reproducible, Data |
 |
2017 |
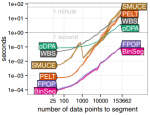
Maidstone R, Hocking T, Rigaill G, Fearnhead P |
On optimal multiple changepoint algorithms for large data |
Statistics and Computing 27 |
Publisher, Software, Reproducible, Local PDF |
 |
2016 |
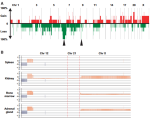
Chicard M, Boyault S, Colmet Daage L, Richer W, Gentien D, Pierron G, Lapouble E, Bellini A, Clement N, Iacono I, Bréjon Sé, Carrere M, Reyes Cé, Hocking T, Bernard V, Peuchmaur M, Corradini Nè, Faure-Conter Cé, Coze C, Plantaz D, Defachelles AS, Thebaud E, Gambart M, Millot Féé, Valteau-Couanet D, Michon J, Puisieux A, Delattre O, Combaret Vé, Schleiermacher G |
Genomic Copy Number Profiling Using Circulating Free Tumor DNA Highlights Heterogeneity in Neuroblastoma |
Clinical Cancer Research 22(22) |
Publisher |
 |
2016 |
Shimada K, Shimada S, Sugimoto K, Nakatochi M, Suguro M, Hirakawa A, Hocking TD, Takeuchi I, Tokunaga T, Takagi Y, Sakamoto A, Aoki T, Naoe T, Nakamura S, Hayakawa F, Seto M, Tomita A, Kiyoi H |
Development and analysis of patient-derived xenograft mouse models in intravascular large B-cell lymphoma |
Leukemia 30(7) |
Pubmed, Local PDF |
 |
2015 |
Hocking TD, Rigaill G, Bourque G |
PeakSeg: constrained optimal segmentation and supervised penalty learning for peak detection in count data |
Proc. 32nd ICML |
Publisher, Video, Software, Reproducible |
 |
2014 |
Hocking TD, Boeva V, Rigaill G, Schleiermacher G, Janoueix-Lerosey I, Delattre O, Richer W, Bourdeaut F, Suguro M, Seto M, Bach F, Vert JP |
SegAnnDB: interactive Web-based genomic segmentation |
Bioinformatics 30(11) |
Pubmed, Software, Reproducible, Preprint, Package, Reproducible |
 |
2014 |
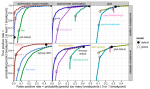
Suguro M, Yoshida N, Umino A, Kato H, Tagawa H, Nakagawa M, Fukuhara N, Karnan S, Takeuchi I, Hocking TD, Arita K, Karube K, Tsuzuki S, Nakamura S, Kinoshita T, Seto M |
Clonal heterogeneity of lymphoid malignancies correlates with poor prognosis |
Cancer Sci 105(7) |
Pubmed |
 |
2013 |
Hocking TD, Schleiermacher G, Janoueix-Lerosey I, Boeva V, Cappo J, Delattre O, Bach F, Vert J-P |
Learning smoothing models of copy number profiles using breakpoint annotations |
BMC Bioinformatics 14(164) |
Publisher, Software, Reproducible |
 |
2013 |
Hocking TD, Wutzler T, Ponting K, Grosjean P |
Sustainable, Extensible Documentation Generation using inlinedocs |
Journal of Statistical Software 54 |
Publisher, Software, Reproducible |
 |
2013 |
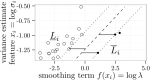
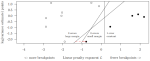
Rigaill G, Hocking T, Vert J-P, Bach F |
Learning sparse penalties for change-point detection using max margin interval regression |
Proc. 30th ICML |
Publisher, Video, Software, Reproducible |
 |
2012 |
Hocking TD |
Learning algorithms and statistical software, with applications to bioinformatics |
PHD thesis, Ecole normale supérieure de Cachan |
Publisher, Reproducible |
 |
2011 |
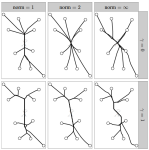
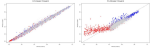
Hocking TD, Joulin A, Bach F, Vert J-P |
Clusterpath an algorithm for clustering using convex fusion penalties |
28th international conference on machine learning |
Publisher, Video, Software, Reproducible, Cited in Ihaka lecture |
 |
2010 |
Gautier M, Hocking TD, Foulley J-L |
A Bayesian outlier criterion to detect SNPs under selection in large data sets |
PLoS one 5(8) |
Publisher |
 |
2008 |
Doyon Y, McCammon JM, Miller JC, Faraji F, Ngo C, Katibah GE, Amora R, Hocking TD, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Amacher SL |
Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases |
Nature biotechnology 26(6) |
Pubmed |











































































